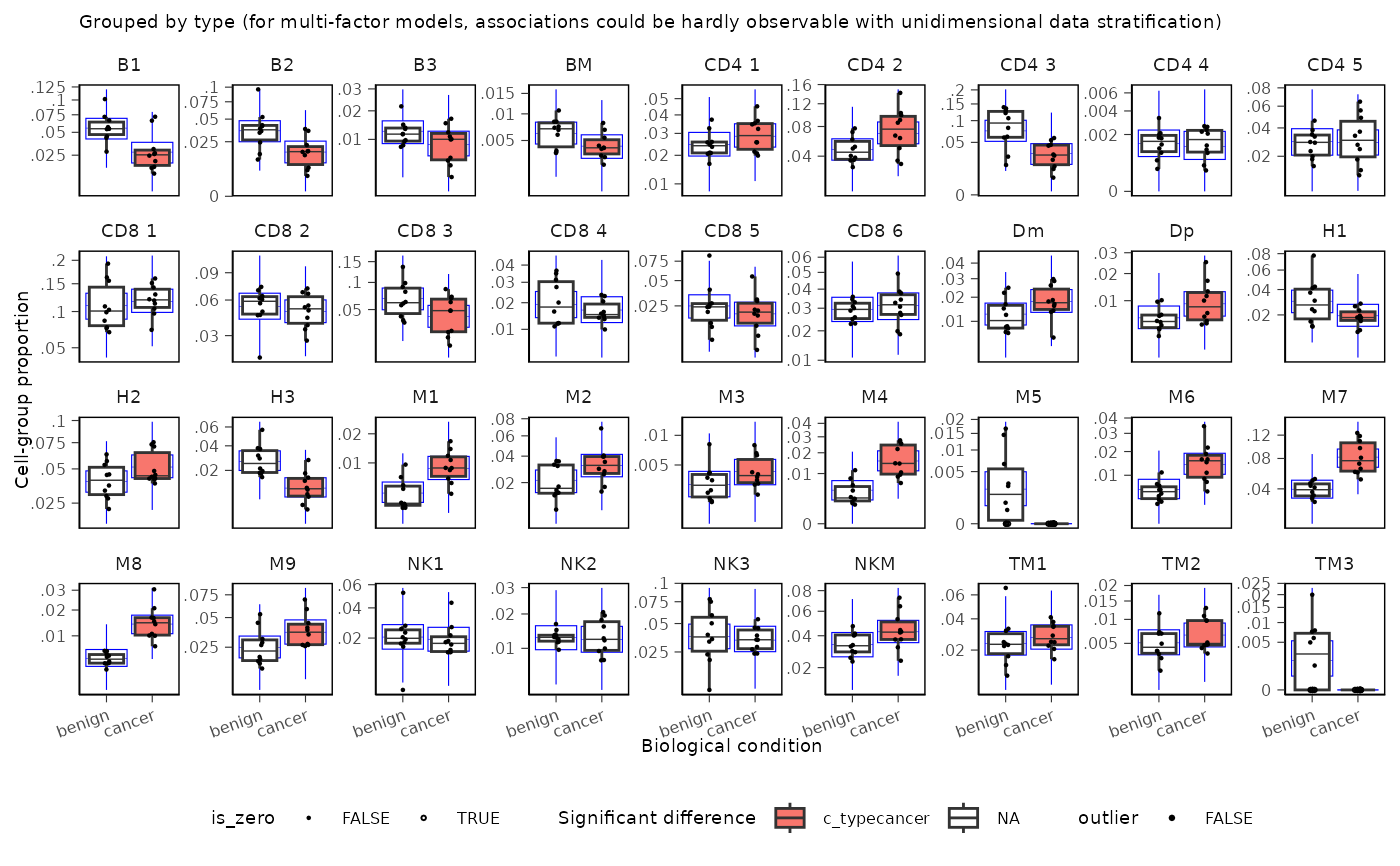
Creates a boxplot visualization of the model results from sccomp. This function plots the estimated cell proportions across samples, highlighting significant changes in cell composition according to a specified factor.
Arguments
- .data
A tibble containing the results from
sccomp_estimateandsccomp_test, including the columns: cell_group name, sample name, read counts, factor(s), p-values, and significance indicators.- factor
A character string specifying the factor of interest included in the model for stratifying the boxplot.
- significance_threshold
A numeric value indicating the threshold for labeling significant cell-groups. Defaults to 0.05.
- significance_statistic
Character vector indicating which statistic is used to colour significant groups. Defaults to
c("pH0", "FDR").- test_composition_above_logit_fold_change
A positive numeric value representing the effect size threshold used in the hypothesis test. A value of 0.2 corresponds to a change in cell proportion of approximately 10% for a cell type with a baseline proportion of 50% (e.g., from 45% to 55%). This threshold is consistent on the logit-unconstrained scale, even when the baseline proportion is close to 0 or 1.
- remove_unwanted_effects
A logical value indicating whether to remove unwanted variation from the data before plotting. Defaults to
FALSE.- cache_stan_model
A character string specifying the cache directory for compiled Stan models. Default is
sccomp_stan_models_cache_dirwhich points to~/.sccomp_models. Use a custom path in restricted environments where the default is not writable.
Value
A ggplot object representing the boxplot of cell proportions across samples, stratified by the specified factor.
References
S. Mangiola, A.J. Roth-Schulze, M. Trussart, E. Zozaya-Valdés, M. Ma, Z. Gao, A.F. Rubin, T.P. Speed, H. Shim, & A.T. Papenfuss, sccomp: Robust differential composition and variability analysis for single-cell data, Proc. Natl. Acad. Sci. U.S.A. 120 (33) e2203828120, https://doi.org/10.1073/pnas.2203828120 (2023).
Examples
print("cmdstanr is needed to run this example.")
#> [1] "cmdstanr is needed to run this example."
# Note: Before running the example, ensure that the 'cmdstanr' package is installed:
# install.packages("cmdstanr", repos = c("https://stan-dev.r-universe.dev/", getOption("repos")))
# \donttest{
if (instantiate::stan_cmdstan_exists()) {
data("counts_obj")
estimate <- sccomp_estimate(
counts_obj,
formula_composition = ~ type,
formula_variability = ~ 1,
sample = "sample",
cell_group = "cell_group",
abundance = "count",
cores = 1
) |>
sccomp_test()
# Plot the boxplot of estimated cell proportions
sccomp_boxplot(
.data = estimate,
factor = "type",
significance_threshold = 0.05
)
}
#> sccomp says: count column is an integer. The sum-constrained beta binomial model will be used
#> sccomp says: estimation
#> sccomp says: the composition design matrix has columns: (Intercept), typecancer
#> sccomp says: the variability design matrix has columns: (Intercept)
#> Loading model from cache...
#> Path [1] :Initial log joint density = -481550.980752
#> Path [1] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 57 -4.788e+05 1.242e-02 2.912e-01 1.000e+00 1.000e+00 3186 -3.686e+03 -3.686e+03
#> Path [1] :Best Iter: [56] ELBO (-3685.745466) evaluations: (3186)
#> Path [2] :Initial log joint density = -481876.692030
#> Path [2] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 53 -4.788e+05 6.763e-03 2.461e-01 7.940e-01 7.940e-01 2951 -3.687e+03 -3.702e+03
#> Path [2] :Best Iter: [51] ELBO (-3686.726337) evaluations: (2951)
#> Path [3] :Initial log joint density = -481670.577303
#> Path [3] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 62 -4.788e+05 8.834e-03 2.118e-01 1.000e+00 1.000e+00 3613 -3.685e+03 -3.697e+03
#> Path [3] :Best Iter: [55] ELBO (-3684.938115) evaluations: (3613)
#> Path [4] :Initial log joint density = -481804.678779
#> Path [4] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 57 -4.788e+05 2.526e-03 2.607e-01 6.946e-01 6.946e-01 3209 -3.687e+03 -3.697e+03
#> Path [4] :Best Iter: [55] ELBO (-3686.573448) evaluations: (3209)
#> Path [5] :Initial log joint density = -481605.905290
#> Path [5] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 9.043e-03 2.310e-01 9.167e-01 9.167e-01 3159 -3.684e+03 -3.691e+03
#> Path [5] :Best Iter: [55] ELBO (-3684.426448) evaluations: (3159)
#> Path [6] :Initial log joint density = -481758.328660
#> Path [6] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 58 -4.788e+05 6.364e-03 2.285e-01 8.408e-01 8.408e-01 3220 -3.690e+03 -3.696e+03
#> Path [6] :Best Iter: [55] ELBO (-3689.697206) evaluations: (3220)
#> Path [7] :Initial log joint density = -481728.012193
#> Path [7] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 5.827e-03 1.931e-01 1.000e+00 1.000e+00 3460 -3.683e+03 -3.685e+03
#> Path [7] :Best Iter: [56] ELBO (-3682.944627) evaluations: (3460)
#> Path [8] :Initial log joint density = -482340.446888
#> Path [8] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 57 -4.788e+05 1.067e-02 3.833e-01 1.000e+00 1.000e+00 3242 -3.684e+03 -3.693e+03
#> Path [8] :Best Iter: [56] ELBO (-3683.783142) evaluations: (3242)
#> Path [9] :Initial log joint density = -481780.453290
#> Path [9] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 57 -4.788e+05 5.278e-03 2.107e-01 1.000e+00 1.000e+00 3151 -3.693e+03 -3.698e+03
#> Path [9] :Best Iter: [55] ELBO (-3693.029372) evaluations: (3151)
#> Path [10] :Initial log joint density = -485608.603964
#> Path [10] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 1.111e-02 2.445e-01 1.000e+00 1.000e+00 3222 -3.693e+03 -3.685e+03
#> Path [10] :Best Iter: [56] ELBO (-3684.631336) evaluations: (3222)
#> Path [11] :Initial log joint density = -481580.668780
#> Path [11] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 1.012e-02 2.591e-01 1.000e+00 1.000e+00 3201 -3.685e+03 -3.688e+03
#> Path [11] :Best Iter: [55] ELBO (-3685.266315) evaluations: (3201)
#> Path [12] :Initial log joint density = -481801.435894
#> Path [12] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 1.070e-02 2.242e-01 1.000e+00 1.000e+00 3415 -3.686e+03 -3.686e+03
#> Path [12] :Best Iter: [56] ELBO (-3685.693177) evaluations: (3415)
#> Path [13] :Initial log joint density = -481484.940634
#> Path [13] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 8.505e-03 1.983e-01 1.000e+00 1.000e+00 3147 -3.686e+03 -3.684e+03
#> Path [13] :Best Iter: [56] ELBO (-3684.053550) evaluations: (3147)
#> Path [14] :Initial log joint density = -481546.869027
#> Path [14] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 9.938e-03 2.185e-01 1.000e+00 1.000e+00 3210 -3.686e+03 -3.685e+03
#> Path [14] :Best Iter: [56] ELBO (-3684.947099) evaluations: (3210)
#> Path [15] :Initial log joint density = -483207.459703
#> Path [15] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 9.285e-03 2.553e-01 9.439e-01 9.439e-01 3108 -3.685e+03 -3.695e+03
#> Path [15] :Best Iter: [55] ELBO (-3685.429071) evaluations: (3108)
#> Path [16] :Initial log joint density = -481563.931669
#> Path [16] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 3.725e-03 2.329e-01 7.556e-01 7.556e-01 3415 -3.683e+03 -3.696e+03
#> Path [16] :Best Iter: [56] ELBO (-3683.188715) evaluations: (3415)
#> Path [17] :Initial log joint density = -481823.269515
#> Path [17] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 61 -4.788e+05 1.153e-02 3.063e-01 1.000e+00 1.000e+00 3551 -3.680e+03 -3.687e+03
#> Path [17] :Best Iter: [60] ELBO (-3679.704731) evaluations: (3551)
#> Path [18] :Initial log joint density = -481937.987153
#> Path [18] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 4.339e-03 2.918e-01 6.817e-01 6.817e-01 3208 -3.689e+03 -3.696e+03
#> Path [18] :Best Iter: [49] ELBO (-3688.685185) evaluations: (3208)
#> Path [19] :Initial log joint density = -482027.282631
#> Path [19] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 57 -4.788e+05 7.523e-03 2.100e-01 1.000e+00 1.000e+00 3241 -3.688e+03 -3.684e+03
#> Path [19] :Best Iter: [57] ELBO (-3683.999957) evaluations: (3241)
#> Path [20] :Initial log joint density = -481663.802218
#> Path [20] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 52 -4.788e+05 1.126e-02 2.352e-01 1.000e+00 1.000e+00 2833 -3.690e+03 -3.699e+03
#> Path [20] :Best Iter: [46] ELBO (-3690.442016) evaluations: (2833)
#> Path [21] :Initial log joint density = -481656.598590
#> Path [21] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 62 -4.788e+05 1.438e-02 4.007e-01 1.000e+00 1.000e+00 3654 -3.683e+03 -3.690e+03
#> Path [21] :Best Iter: [61] ELBO (-3683.292788) evaluations: (3654)
#> Path [22] :Initial log joint density = -481598.542624
#> Path [22] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 55 -4.788e+05 7.161e-03 1.684e-01 1.000e+00 1.000e+00 3026 -3.690e+03 -3.686e+03
#> Path [22] :Best Iter: [55] ELBO (-3686.168912) evaluations: (3026)
#> Path [23] :Initial log joint density = -481997.811304
#> Path [23] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 8.092e-03 2.580e-01 1.000e+00 1.000e+00 3305 -3.684e+03 -3.686e+03
#> Path [23] :Best Iter: [56] ELBO (-3684.259783) evaluations: (3305)
#> Path [24] :Initial log joint density = -482171.664488
#> Path [24] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 62 -4.788e+05 1.378e-02 3.663e-01 1.000e+00 1.000e+00 3713 -3.683e+03 -3.692e+03
#> Path [24] :Best Iter: [59] ELBO (-3683.305495) evaluations: (3713)
#> Path [25] :Initial log joint density = -483386.401310
#> Path [25] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 52 -4.788e+05 5.627e-03 1.862e-01 1.000e+00 1.000e+00 2875 -3.691e+03 -3.696e+03
#> Path [25] :Best Iter: [45] ELBO (-3690.983033) evaluations: (2875)
#> Path [26] :Initial log joint density = -481637.258691
#> Path [26] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 61 -4.788e+05 2.097e-02 2.013e-01 1.000e+00 1.000e+00 3606 -3.683e+03 -3.682e+03
#> Path [26] :Best Iter: [61] ELBO (-3682.027363) evaluations: (3606)
#> Path [27] :Initial log joint density = -481618.067917
#> Path [27] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 1.079e-02 2.681e-01 8.899e-01 8.899e-01 3305 -3.683e+03 -3.696e+03
#> Path [27] :Best Iter: [57] ELBO (-3682.889239) evaluations: (3305)
#> Path [28] :Initial log joint density = -482398.942985
#> Path [28] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 58 -4.788e+05 8.023e-03 2.000e-01 9.497e-01 9.497e-01 3277 -3.683e+03 -3.692e+03
#> Path [28] :Best Iter: [56] ELBO (-3682.798658) evaluations: (3277)
#> Path [29] :Initial log joint density = -481498.760395
#> Path [29] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 1.286e-02 2.701e-01 1.000e+00 1.000e+00 3086 -3.686e+03 -3.683e+03
#> Path [29] :Best Iter: [56] ELBO (-3683.433002) evaluations: (3086)
#> Path [30] :Initial log joint density = -481539.223694
#> Path [30] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 1.059e-02 1.836e-01 1.000e+00 1.000e+00 3390 -3.684e+03 -3.683e+03
#> Path [30] :Best Iter: [59] ELBO (-3682.914744) evaluations: (3390)
#> Path [31] :Initial log joint density = -485405.720368
#> Path [31] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 52 -4.788e+05 4.562e-03 2.059e-01 1.000e+00 1.000e+00 2976 -3.692e+03 -3.708e+03
#> Path [31] :Best Iter: [40] ELBO (-3691.752195) evaluations: (2976)
#> Path [32] :Initial log joint density = -481767.625615
#> Path [32] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 55 -4.788e+05 1.029e-02 1.661e-01 1.000e+00 1.000e+00 3118 -3.693e+03 -3.683e+03
#> Path [32] :Best Iter: [55] ELBO (-3682.837103) evaluations: (3118)
#> Path [33] :Initial log joint density = -481589.553067
#> Path [33] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 8.866e-03 2.066e-01 1.000e+00 1.000e+00 3305 -3.682e+03 -3.684e+03
#> Path [33] :Best Iter: [58] ELBO (-3682.005192) evaluations: (3305)
#> Path [34] :Initial log joint density = -481767.585540
#> Path [34] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 7.548e-03 2.480e-01 1.000e+00 1.000e+00 3252 -3.687e+03 -3.693e+03
#> Path [34] :Best Iter: [53] ELBO (-3687.438710) evaluations: (3252)
#> Path [35] :Initial log joint density = -481704.945037
#> Path [35] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 58 -4.788e+05 7.448e-03 2.091e-01 8.174e-01 8.174e-01 3383 -3.681e+03 -3.694e+03
#> Path [35] :Best Iter: [56] ELBO (-3681.356368) evaluations: (3383)
#> Path [36] :Initial log joint density = -481643.606635
#> Path [36] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 1.074e-02 3.983e-01 1.000e+00 1.000e+00 3402 -3.683e+03 -3.693e+03
#> Path [36] :Best Iter: [56] ELBO (-3683.237489) evaluations: (3402)
#> Path [37] :Initial log joint density = -483368.973003
#> Path [37] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 57 -4.788e+05 9.690e-03 3.549e-01 9.692e-01 9.692e-01 3241 -3.684e+03 -3.691e+03
#> Path [37] :Best Iter: [55] ELBO (-3683.591942) evaluations: (3241)
#> Path [38] :Initial log joint density = -481832.246063
#> Path [38] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 55 -4.788e+05 1.609e-02 2.477e-01 1.000e+00 1.000e+00 3133 -3.692e+03 -3.685e+03
#> Path [38] :Best Iter: [55] ELBO (-3685.014173) evaluations: (3133)
#> Path [39] :Initial log joint density = -483031.934991
#> Path [39] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 57 -4.788e+05 8.618e-03 3.057e-01 9.450e-01 9.450e-01 3405 -3.686e+03 -3.692e+03
#> Path [39] :Best Iter: [55] ELBO (-3686.367500) evaluations: (3405)
#> Path [40] :Initial log joint density = -481705.412677
#> Path [40] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 58 -4.788e+05 3.554e-03 2.568e-01 7.233e-01 7.233e-01 3358 -3.684e+03 -3.695e+03
#> Path [40] :Best Iter: [55] ELBO (-3683.688259) evaluations: (3358)
#> Path [41] :Initial log joint density = -484236.817694
#> Path [41] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 59 -4.788e+05 5.572e-03 2.417e-01 1.000e+00 1.000e+00 3352 -3.686e+03 -3.688e+03
#> Path [41] :Best Iter: [55] ELBO (-3685.671263) evaluations: (3352)
#> Path [42] :Initial log joint density = -481602.782678
#> Path [42] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 56 -4.788e+05 3.007e-03 1.765e-01 7.398e-01 7.398e-01 3190 -3.691e+03 -3.696e+03
#> Path [42] :Best Iter: [46] ELBO (-3690.649952) evaluations: (3190)
#> Path [43] :Initial log joint density = -484710.001137
#> Path [43] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 61 -4.788e+05 3.810e-03 2.298e-01 7.213e-01 7.213e-01 3633 -3.685e+03 -3.696e+03
#> Path [43] :Best Iter: [55] ELBO (-3684.766956) evaluations: (3633)
#> Path [44] :Initial log joint density = -481724.225258
#> Path [44] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 51 -4.788e+05 3.816e-03 2.414e-01 8.410e-01 8.410e-01 2992 -3.688e+03 -3.703e+03
#> Path [44] :Best Iter: [43] ELBO (-3688.281245) evaluations: (2992)
#> Path [45] :Initial log joint density = -481538.037530
#> Path [45] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 53 -4.788e+05 5.494e-03 2.173e-01 9.411e-01 9.411e-01 2869 -3.690e+03 -3.702e+03
#> Path [45] :Best Iter: [52] ELBO (-3690.187879) evaluations: (2869)
#> Path [46] :Initial log joint density = -481690.773823
#> Path [46] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 62 -4.788e+05 1.325e-02 1.829e-01 1.000e+00 1.000e+00 3734 -3.683e+03 -3.680e+03
#> Path [46] :Best Iter: [62] ELBO (-3680.388405) evaluations: (3734)
#> Path [47] :Initial log joint density = -481601.061523
#> Path [47] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 58 -4.788e+05 7.424e-03 2.257e-01 1.000e+00 1.000e+00 3220 -3.682e+03 -3.690e+03
#> Path [47] :Best Iter: [55] ELBO (-3681.716140) evaluations: (3220)
#> Path [48] :Initial log joint density = -481849.557099
#> Path [48] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 65 -4.788e+05 1.161e-02 2.703e-01 1.000e+00 1.000e+00 3889 -3.683e+03 -3.682e+03
#> Path [48] :Best Iter: [65] ELBO (-3682.253048) evaluations: (3889)
#> Path [49] :Initial log joint density = -481572.243009
#> Path [49] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 60 -4.788e+05 8.202e-03 1.962e-01 1.000e+00 1.000e+00 3422 -3.684e+03 -3.686e+03
#> Path [49] :Best Iter: [58] ELBO (-3683.770019) evaluations: (3422)
#> Path [50] :Initial log joint density = -482693.043960
#> Path [50] : Iter log prob ||dx|| ||grad|| alpha alpha0 # evals ELBO Best ELBO Notes
#> 58 -4.788e+05 5.893e-03 1.842e-01 1.000e+00 1.000e+00 3295 -3.684e+03 -3.684e+03
#> Path [50] :Best Iter: [55] ELBO (-3683.932807) evaluations: (3295)
#> Finished in 13.6 seconds.
#> sccomp says: to do hypothesis testing run `sccomp_test()`,
#> the `test_composition_above_logit_fold_change` = 0.1 equates to a change of ~10%, and
#> 0.7 equates to ~100% increase, if the baseline is ~0.1 proportion.
#> Use `sccomp_proportional_fold_change` to convert c_effect (linear) to proportion difference (non-linear).
#> sccomp says: auto-cleanup removed 1 draw files from 'sccomp_draws_files'
#> sccomp says: When visualising proportions, especially for complex models, consider setting `remove_unwanted_effects=TRUE`. This will adjust the proportions, preserving only the observed effect.
#> sccomp says: from version 2.1.25, the default `significance_statistic` for boxplots is `pH0` (previously `FDR`). Set `significance_statistic = "FDR"` to use the previous default.
#> Precompiled model not found. Compiling the model...
#> Running make /tmp/Rtmp3B3vvJ/model-3b833b775517 "STAN_THREADS=TRUE" \
#> "STANCFLAGS += --include-paths=/tmp/Rtmp3B3vvJ/temp_libpath3b8342d90e1b/sccomp/stan --name='glm_multi_beta_binomial_generate_data_model'"
#>
#> --- Translating Stan model to C++ code ---
#> bin/stanc --include-paths=/tmp/Rtmp3B3vvJ/temp_libpath3b8342d90e1b/sccomp/stan --name='glm_multi_beta_binomial_generate_data_model' --o=/tmp/Rtmp3B3vvJ/model-3b833b775517.hpp /tmp/Rtmp3B3vvJ/model-3b833b775517.stan
#>
#> --- Compiling C++ code ---
#> g++ -Wno-deprecated-declarations -std=c++17 -pthread -D_REENTRANT -Wno-sign-compare -Wno-ignored-attributes -Wno-class-memaccess -DSTAN_THREADS -I stan/lib/stan_math/lib/tbb_2020.3/include -O3 -I src -I stan/src -I stan/lib/rapidjson_1.1.0/ -I lib/CLI11-1.9.1/ -I stan/lib/stan_math/ -I stan/lib/stan_math/lib/eigen_3.4.0 -I stan/lib/stan_math/lib/boost_1.87.0 -I stan/lib/stan_math/lib/sundials_6.1.1/include -I stan/lib/stan_math/lib/sundials_6.1.1/src/sundials -DBOOST_DISABLE_ASSERTS -c -Wno-ignored-attributes -x c++ -o /tmp/Rtmp3B3vvJ/model-3b833b775517.o /tmp/Rtmp3B3vvJ/model-3b833b775517.hpp
#>
#> --- Linking model ---
#> g++ -Wno-deprecated-declarations -std=c++17 -pthread -D_REENTRANT -Wno-sign-compare -Wno-ignored-attributes -Wno-class-memaccess -DSTAN_THREADS -I stan/lib/stan_math/lib/tbb_2020.3/include -O3 -I src -I stan/src -I stan/lib/rapidjson_1.1.0/ -I lib/CLI11-1.9.1/ -I stan/lib/stan_math/ -I stan/lib/stan_math/lib/eigen_3.4.0 -I stan/lib/stan_math/lib/boost_1.87.0 -I stan/lib/stan_math/lib/sundials_6.1.1/include -I stan/lib/stan_math/lib/sundials_6.1.1/src/sundials -DBOOST_DISABLE_ASSERTS -Wl,-L,"/home/runner/.cmdstan/cmdstan-2.38.0/stan/lib/stan_math/lib/tbb" -Wl,-rpath,"/home/runner/.cmdstan/cmdstan-2.38.0/stan/lib/stan_math/lib/tbb" /tmp/Rtmp3B3vvJ/model-3b833b775517.o src/cmdstan/main_threads.o -ltbb stan/lib/stan_math/lib/sundials_6.1.1/lib/libsundials_nvecserial.a stan/lib/stan_math/lib/sundials_6.1.1/lib/libsundials_cvodes.a stan/lib/stan_math/lib/sundials_6.1.1/lib/libsundials_idas.a stan/lib/stan_math/lib/sundials_6.1.1/lib/libsundials_kinsol.a stan/lib/stan_math/lib/tbb/libtbb.so.2 -o /tmp/Rtmp3B3vvJ/model-3b833b775517
#> rm /tmp/Rtmp3B3vvJ/model-3b833b775517.o /tmp/Rtmp3B3vvJ/model-3b833b775517.hpp
#> Model compiled and saved to cache successfully.
#> Running standalone generated quantities after 1 MCMC chain, with 1 thread(s) per chain...
#>
#> Chain 1 finished in 0.0 seconds.
#> Joining with `by = join_by(cell_group, sample)`
#> Joining with `by = join_by(cell_group, type)`
 # }
# }